Control of Gene Expression
All cells contain a set of genes, which can be thought of as a set of instructions for making each of a very large number of proteins . The creation of a protein from its gene is called gene expression. However, for a given cell not all of these instructions are actually used, and among those that are, some are used more than others or only under certain circumstances. Controlling gene expression is critical to a cell because it allows it to avoid wasting energy and raw materials in the synthesis of proteins it does not need. Thus, it allows a cell to be a more streamlined and versatile entity that can respond to changing conditions by adjusting its physiology.
To understand the control of gene expression, two key concepts should be understood. First, gene expression requires transcription , the process of making a messenger ribonucleic acid (mRNA) copy of the deoxyribonucleic acid (DNA) gene. Transcription can only occur if RNA polymerase first attaches, or binds, to the DNA. Controlling this binding process is the major way that gene expression is controlled, and proteins are the major controllers of binding.
The second important concept is that a protein molecule that helps regulate binding can itself be regulated. This usually occurs when some other molecule binds to the protein, causing the protein to undergo a structural change, in other words, to change shape. In some cases this shape change will help RNA polymerase to bind to DNA, and in other cases it will prevent it from doing so.
Control in Prokaryotes
Negative Control. The concept that gene expression could be controlled originated with studies done in the 1950s by French scientists François Jacob and Jacques Monod. They were studying the metabolism of a sugar, called lactose, by the E. coli bacterium. β-Lactose metabolism requires three proteins. Galactosidase and lactose permease are both involved directly in lactose metabolism; β-galactosidase hydrolyzes lactose into galactose and glucose , and lactose permease transports lactose across the bacterial cell membrane. The physiologic role of the third protein, thiogalactoside acetylase, is unclear. Jacob and Monod found that the amount of the three proteins all increased when E. coli were cultured in lactose-containing medium (a nutrient source). This led to the hypothesis that the three genes were regulated together as a single unit.
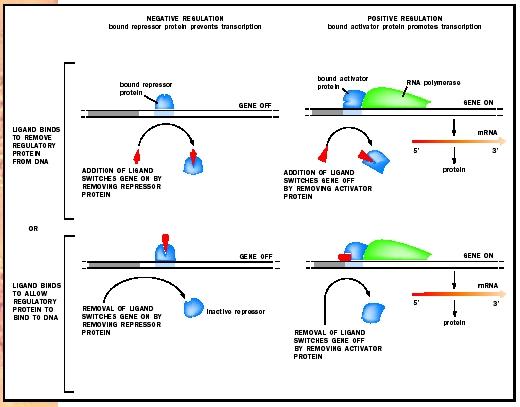
This type of multigene unit was dubbed an "operon" and consists of the structural genes, which encode proteins, plus regulatory sequences lying upstream on the DNA. The structural genes in an operon are transcribed as a single mRNA, and the mRNA is thus polygenic (or polycistronic). An elegant series of experiments showed that transcription was begun when a lactose derivative, Allolactose, caused a repressor to be removed from the transcription initiation site. Thus, lactose regulates the synthesis of the enzymes necessary for its own metabolism by releasing the transcriptional repression imposed upon them. This type of regulation is called negative regulation, since it employs a repression to prevent transcription. The use of activator proteins in the positive control of gene expression is also common in prokaryotes . In this system, the activator protein promotes transcription.
Positive Control. Positive control of gene expression is illustrated by the transcriptional activator, catabolite gene activator protein (CAP). CAP activates transcription of the lac operon, in addition to many other inducible operons. Because glucose is a preferred food source, the lac operon is not activated in E. coli cells cultured in medium containing both glucose and lactose until the glucose is used up. However, since lactose is present, one might expect the lac operon to be derepressed and hence active. But experiments have shown that glucose itself represses the activity of the lac operon, such that only when lactose is the only source of energy is it activated. This glucose repression is observed for a number of other operons that encode enzymes for the utilization of alternative energy sources. Glucose repression occurs via a positive mechanism. As glucose is consumed, its level in the cell drops. Low glucose levels stimulate the production of a small molecule called cyclic- AMP (cAMP), which then binds CAP. CAP undergoes a structural change that allows it to bind DNA and activate transcription. Thus, regulation of the lac operon is achieved by a collaboration between the negative control of the lac repressor and the positive control of CAP.
The lac repressor and CAP are examples of regulators of initiation of transcription. Although most regulators act at this level, some act at the level of elongation of the mRNA, after transcription has started. The tryptophan operon (trp operon) consists of five structural genes necessary for the biosynthesis of the amino acid tryptophan. It is regulated at the level of initiation via a negative regulatory scheme much like that for the lac operon; however, an additional mechanism, called transcriptional attenuation , is also at work. Part of the mRNA generated from the trp operon spontaneously folds into a stem-loop structure that exposes a termination sequence, causing transcription to terminate prematurely. However, when tryptophan is lacking, the ribosome works more slowly (since tryptophan is needed to make protein). This allows time for the formation of a different structure, the stem-loop, which hides the termination sequence, with the result being that transcription continues and a full-length transcript is produced. Thus, the end product of the operon, tryptophan, actively participates in the regulation of its own synthesis. This is a common theme in prokaryotic transcriptional regulation. Transcriptional attenuation can occur in prokaryotes
Eukaryotic Transcription
Regulation of transcription is by necessity far more complex in eukaryotic cells (cells with a nucleus ) than in prokaryotic cells. Not only are eukaryotic cells larger and more highly compartmentalized, but multicellular eukaryotes pass through a number of developmental stages, each requiring different proteins, on the road to their final differentiated state. Also, multicellular organisms contain many different cell types, each of which expresses distinct sets of proteins.
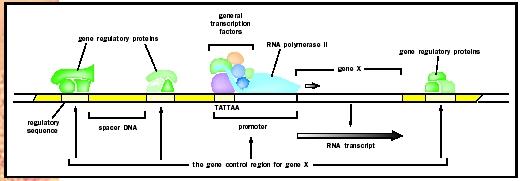
Certain basic features of transcriptional regulation are shared between prokaryotes and eukaryotes; in both cases it involves an interplay between activators and repressors that bind cis-acting sequences on DNA. However, one major difference is that, unlike prokaryotic DNA, eukaryotic chromosomes are wrapped around proteins called histones , to form a condensed form of DNA called chromatin . This tends to repress gene transcription, and several transcriptional activators have been found to function by relieving chromatin-induced repression. Another feature that distinguishes eukaryotic from prokaryotic transcription is that RNA polymerase does not bind directly to DNA but instead binds via a set of proteins called the basic transcription factor . Thus, in many cases the role of activators is to recruit these transcription factors to the promoter site rather than to directly recruit the polymerase itself. Finally, whereas prokaryotic genes are often controlled by only one or two regulatory proteins, eukaryotic genes are typically controlled by a multiplicity of factors. This added complexity allows for the fine-tuning of gene activity in response to multiple stimuli.
Structure of Transcriptional Activators
Many transcriptional activators are essentially modular in structure in that the DNA-binding domain and the transactivation (or activation) domain can almost be thought of as two distinct proteins that are physically linked. The DNA-binding domain is the part of the molecule that contacts DNA at the promoter site. The transactivation domain is the part that recruits other factors to the promoter such that the rate of transcription of the gene increases. Although transcription factor DNA-binding domains vary in amino acid sequence, many can be placed into structural categories based on their three-dimensional structures. Among these are the zinc finger, helix-loop-helix,
Regulation of Transcriptional Activators
Regulation of transcription sometimes occurs via the simple presence or absence of transcription factors. An example of this is in the regulation of the immunoglobulin (an immune protein, also called antibody ) heavy chain gene, which is expressed in B lymphocytes (white blood cells that make antibodies) but not other cell types. This gene's enhancer (a region distant from the promoter) contains at least nine binding sites for regulatory proteins. The enhancer is acted on by activators present in B lymphocytes , while in nonlymphocyte cells repressors are present that inhibit transcription. This limits expression of the gene to lymphocytes.
Often, however, regulation does not occur at the level of presence or absence of a regulatory protein but rather by modulation of its activity. Thus, many transcription factors are always present in the cell, awaiting the specific signals that will convert them from an inactive to an active form. How is this achieved? The three most common mechanisms are regulation of nuclear localization, regulation of DNA binding, and regulation of transactivation.
Regulation of Nuclear Localization. In many cases a protein is kept in the cytoplasm , well away from its target genes, until a stimulus signals it to enter the nucleus and activate transcription. This mode of regulation works because transport into the nucleus is regulated, such that only proteins possessing a special tag are allowed to enter. The transcription factor NF-κB, which regulates a number of genes in immune cells that help fight infections, is regulated in this way. NF-κB is present in the cytoplasm of unstimulated immune cells as a complex with an inhibitory protein called iκB. Upon receiving a stimulus, such as a viral infection, iκB becomes phosphorylated and is subsequently degraded, leaving NF-κB free to enter the nucleus and activate its target genes to help fight infection. Interestingly, one of these target genes is the i B gene, and thus inhibition of NF-κB is reestablished shortly thereafter. This kind of negative feedback mechanism, bringing the cell back to its unstimulated state, is common among inducible genes. In addition, NF-κB activation illustrates another common feature of transcription factor regulation in eukaryotes: phosphorylation is often used as a switch that interconverts a transcription factor back and forth between inactive and active forms.
Regulation of DNA Binding. A second common mechanism by which the activity of a transcription factor is controlled is through alteration of its DNA-binding ability. The steroid hormone receptor family is a good example of this. This family of transcription factors has many members, all related in structure, yet binding to distinct steroid hormones on the one hand, and activating distinct sets of genes on the other. Some of these hormone receptors reside in the cytoplasm and others in the nucleus, but all are unable to bind their target DNA sequence until they first bind to their corresponding steroid hormone. This causes them to undergo a conformational change that increases their affinity for DNA, allowing them to bind. It is through their action on hormone receptors and DNA that steroid hormones exert their powerful effects on the body's cells.
Another way to increase the DNA-binding ability of a transcription factor is to induce it to multimerize. Many factors are inactive by themselves, but when induced to bind other factors, they can bind their target sequences and activate transcription. The other factors can either be identical molecules of the same factor, thus forming homo-multimers, or different proteins, forming hetero-multimers. An example of this occurs with heat shock factor (HSF) in mammalian cells, which upon stimulation forms homotrimers. The DNA-binding affinity of a single molecule of HSF for its binding site is too low to be physiologically significant; however, a complex of three molecules binds the target site very tightly, making HSF one of the most inducible transcription factors known.
Regulation of Transactivation. Finally, some transcriptional activators are already bound to their target sites in gene promoters but remain transcriptionally inactive until they are stimulated. In yeast, HSF is already trimerized and bound to some of its target genes in unstimulated cells. Heat shock (a rise in temperature) results in phosphorylation of HSF at multiple sites, which induces a structural change in the protein that unleashes the transactivation domain.
The aforementioned examples illustrate a number of ways in which a transcriptional activator may be regulated. However, it should be kept in mind that many are regulated in more than one way. For example, both nuclear localization and DNA-binding ability of an activator may be controlled. Thus, even if a few molecules should happen into the nucleus by mistake, they would not be able to bind and activate their target genes. This kind of tight control is important because sometimes even small levels of a protein can set off a cascade of reactions that can dramatically change the physiology of the cell. It is critical to avoid these types of false alarms in order for the cell not to waste valuable energy and resources, and so that it remains poised to respond to a genuine stimulus.
Transcriptional Repression
Transcriptional repressors, like activators, bind cis-acting sequences in the genes they regulate and are modular in structure, possessing distinct DNA-binding and repressor domains. However, as their name implies, their role is in the repression of gene activity rather than their activation. Some repressors function by simply binding upstream regions of genes and blocking the binding of either activator proteins or the polymerase itself, much like the repressor in the lac operon. Some extremely versatile proteins can function either as repressors or activators, depending on the proteins with which they interact. An example is the Mcm1 protein in yeast. Yeast can be one of two mating types, called α and ("alpha"), each of which expresses mating type-specific sets of genes. Mcm1 dimerizes with one protein to repress the a-specific genes in α cells, and with another to activate the α-specific genes.
The Role of Chromatin
Although transcriptional repressors often participate in gene regulation, it must be kept in mind that the very nature of DNA in eukaryotic cells tends to keep genes in the repressed state. Eukaryotic DNA is wrapped around protein complexes called histone octamers, which has the effect of packaging the DNA into a compact form such that it fits inside the nucleus. However, this also limits access of regulatory factors to their target sites. As the mechanisms of transcriptional activators are being uncovered, more and more are being found that act by relieving chromatin-induced repression. An example is the Swi/Snf protein complex, first identified in yeast. Mutations in components of the complex resulted in decreased activity of certain target genes. It was later found that mutations in the histone genes restored normal activity to those target genes; in other words, the mutations in the histone genes somehow compensated for the mutations in Swi/Snf. This was an indication that histones and Swi/Snf interact in some way and suggested that Swi/Snf might function by disrupting histone binding to DNA. Biochemical experiments carried out later on showed that this was indeed the case. Although Swi/Snf does not completely dissociate histones from DNA, it loosens them, which is sufficient to allow many activators to bind. Swi/Snf is only involved in activating a subset of genes, and the question of why it functions at some promoters and not others is a topic of intense research.
A second mechanism by which chromatin-induced repression is relieved is by histone acetylation . Histones are positively charged proteins and hence interact tightly with DNA, which is negatively charged. Acetylation of histones reduces their net positive charge, which loosens their interaction with DNA and increases transcription factor binding. Several transcription factors in a variety of organisms have now been found to be acetyltransferases; in effect, they can acetylate histones.
In addition, some transcriptional repressors in yeast and mammals have been found to be histone deacetylases. In fact, the protein MeCP2, which binds to methylated DNA, has been found to function in a complex with a histone deacetylase. Thus, methylation would lead to binding of this complex, causing deacetylation of histones and a more condensed chromatin structure. Methylated DNA has long been known to be associated with transcriptionally inactive genes, and inroads into the study of histone acetylation have finally provided an explanation for this.
SEE ALSO Chromosome, Eukaryotic ; Control Mechanisms ; DNA ; Gene
Kirstie Saltsman
Bibliography
Stryer, Lubert. "Control of Gene Expression in Prokaryotes." In Biochemistry, 4th ed. New York: W. H. Freeman and Company, 1995.
——. "Eukaryotic Chromosomes and Gene Expression." In Biochemistry, 4th ed. New York: W. H. Freeman and Company, 1995.
REGULATION OF THE LAC OPERON
E. coli with defects in the regulation of the lac operon were found to have mutations in one of two loci, called o and i, located upstream of the structural genes. Mutations in o yielded cells that constitutively (continually) expressed the lac operon, whereas mutations in i fell into two categories; one in which the lac operon was constitutively expressed, and the other in which it was uninducible (could not be expressed). Subsequent experiments showed that i was a gene for a diffusible protein that was the repressor of the lac operon, whereas o was a DNA sequence to which a repressor bound.
This was consistent with the mutant results: A mutation in o would disrupt the binding of the repressor protein, leading to constitutive expression of the lac operon, and a mutation in i would either prevent the repressor from binding to o, resulting in constitutive activation, or render the repressor unresponsive to the inducer, lactose, which would cause uninducibility. Because i was diffusible (could move within the cell) and could interact with any piece of DNA containing its target sequence, it was called a trans-acting factor ( trans means "across"). In contrast, o only affects the genes to which it is physically linked and so has been called a cis-acting factor ( cis means "together"). These elegant genetic studies paved the way for biochemical studies carried out in the 1960s by Walter Gilbert and Benno Müller-Hill. They purified the lac repressor, encoded by i, and found that it bound to a 30 base-pair region of DNA spanning the transcription initiation site, consistent with the location of o. In addition, they found that the lac repressor released its hold on o when bound to allolactose, a derivative of lactose.
In the laboratory the DNA-binding domain and the transactivation domain—the two functional domains—can be mixed and matched between different transcription factors to yield hybrid molecules that still function, albeit differently from the original proteins. This feature has been exploited experimentally. For example, the relative strengths of various activation domains can be assessed by fusing each to the same DNA-binding domain and determining the rate at which each promotes transcription.
Mutations in the MeCP2 cause Rett syndrome, an X-linked dominant disorder marked by seizures, abnormal movements, and mutism.
Comment about this article, ask questions, or add new information about this topic: