Electrophoresis

Electrophoresis is one of the most important techniques used by molecular biologists. To name only a few applications, deoxyribonucleic acid (DNA) electrophoresis is used to map the order of restriction fragments within chromosomes , to analyze DNA variation within a population by restriction fragment length polymorphisms (RFLPs), and to determine the nucleotide sequence of a piece of DNA.
Electrophoresis refers to the migration of a charged molecule through a restrictive matrix , or gel, drawn by an electrical force. As the force drags the molecule through the gel, it encounters resistance from the strands of the gel, retarding its rate of migration. In gel electrophoresis, larger molecules migrate more slowly than smaller ones, and so the distance of migration within a gel can be used to determine a molecule's size.
Although it is possible to separate whole chromosomes using specialized electrophoresis techniques, DNA that is to be analyzed by electrophoresis is usually cut into smaller pieces using restriction enzymes . Fragments of DNA prepared by treatment with restriction enzymes are commonly separated from one another, and their sizes determined, using a gel of agarose electrophoresis, a complex carbohydrate . DNA is negatively charged due to the phosphodiester bonds that join the individual nucleotide building blocks. DNA will therefore electrophorese toward the positive electrode when placed in an electrical field. To visualize the results after electrophoresis, the gel is soaked in a solution that causes DNA to fluoresce when exposed to ultraviolet light.
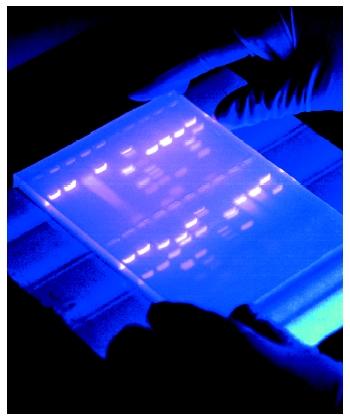
Treatment of the DNA sample with multiple restriction enzymes in various combinations enables the researcher to generate a restriction map of the original DNA fragment, which identifies the sites at the DNA where the restriction enzymes are.
Many research questions require a detailed analysis of one specific DNA fragment in a complex mixture. In such cases, a radioactive DNA probe can be used to identify the fragment based on its nucleotide sequence. The method, known as hybridization, is based on the rules of complementary base pairing (A bonds to T, G bonds to C). A probe is designed whose sequence is complementary to the piece of DNA to be detected. The gel-separated DNA is first transferred to a nylon membrane using a technique called a Southern blot.
During the blotting procedure, the strands within the DNA double helix are separated from each other, or denatured, by treatment with a base. Because double-stranded DNA is more stable than single-stranded, during the hybridization the single-stranded probe will locate and bind to the single-stranded gel-separated fragment with complementary sequence. Being fluorescent or radioactive, the position of the probe can be determined using photographic methods. The target sequence can then be removed by cutting at the piece of the gel that contains it.
The most common technique for determining DNA sequence is the Sanger method, which generates fragments that differ in length by a single nucleotide. High-resolution polyacrylamide gel electrophoresis is then used to separate the fragments and to allow the sequence to be determined.
Electrophoresis of ribonucleic acid (RNA) is an integral procedure in many studies of gene expression . RNA is isolated, separated by electrophoresis, and then the gel-separated RNA fragments are transferred to a nylon membrane using a technique called a Northern blot. Hybridization with a single-stranded DNA probe is then used to determine the position of a specific RNA fragment.
DNA and RNA are relatively simple in terms of structure and composition. Proteins , however, are composed of twenty different amino acids in various combinations, and proteins vary significantly in their three-dimensional structure. The composition of amino acids will affect the charge on the protein, which ultimately will affect its electrophoretic behavior. The shape of a protein similarly will affect its rate of migration. As a result, a specialized technique, SDS-polyacrylamide gel electrophoresis (SDS-PAGE), is usually used to analyze proteins. In this method, protein samples are heated and then treated with the detergent sodium dodecyl sulfate (SDS). Proteins treated in this way are unfolded, linear, and uniformly coated by negatively charged detergent molecules. The rate of migration of treated proteins is inversely proportional to the logarithm of molecular weight. Following electrophoresis, the protein in the gel can be stained to visualize all the proteins in a sample, or the proteins in the gel can be transferred to a nylon membrane (Western blot) and specific ones detected with the use of enzyme -linked antibodies.
Regardless of the macromolecule being studied, gel electrophoresis is a crucial technique to the molecular biologist. Many scientific questions can be answered using electrophoresis, and as a result an active molecular biology research lab will have several benches that are devoted to the required specialized reagents and equipment.
SEE ALSO DNA Sequencing
James E. Blankenship
Bibliography
Alberts, Bruce, et al. Molecular Biology of the Cell, 4th ed. New York: Garland Publishing, 2000.
Stryer, Lubert. Biochemistry, 4th ed. New York: W. H. Freeman and Company, 1995.
The rate of migration is inversely proportional to the logarithm of a molecule's size.
Agarose, which is used to make electrophoresis gel, is derived from the seaweed agar.
SANGER, FREDERICK (1918–)
English biochemist who received two Nobel Prizes in chemistry. The first came in 1958, for finding the amino sequence of insulin, the protein that helps regulate blood sugar levels, and the second, in 1980, for inventing a technique to sequence the nucleotides in a strand of deoxyribonucleic acid (DNA).
Comment about this article, ask questions, or add new information about this topic: