Genetic Code

The genetic code allows an organism to translate the genetic information found in its chromosomes into usable proteins . Stretches of deoxyribonucleic acid (DNA) are built from four different nucleotide bases, while proteins are made from twenty unique subunits called amino acids . This numerical disparity presents an interesting problem: How does the cell translate the genetic information in the four-letter alphabet of DNA into the twenty-letter alphabet of protein? The conversion code is called the genetic code.
Requirements of a Code
The information transfer from DNA to protein, called gene expression , occurs in two steps. In the first step, called transcription , a DNA sequence is copied to make a template for protein synthesis called messenger ribonucleic acid (messenger RNA, or mRNA). During protein synthesis, ribosomes and transfer RNA (tRNA) use the genetic code to convert genetic information contained in mRNA into functional protein. (Formally speaking, the genetic code refers to the RNA-amino acid conversion code and not to DNA, though usage has expanded to refer more broadly to DNA.)
Mathematics reveals the minimum requirements for a genetic code. The ribosome must convert mRNA sequences that are written in four bases—A, G, U, and C—into proteins, which are made up of twenty different amino acids. A one base to one amino acid correspondence would code for only four amino acids (4 1 ). Similarly, all combinations of a two-base code (for example, AA, AU, AG, AC, etc.) will provide for only sixteen amino acids (4 2 ). However, blocks of three RNA bases allow sixty-four (4 3 ) combinations of the four nucleotides, which is more than enough combinations to correspond to the twenty distinct amino acids. So, the genetic code must use blocks of at least three RNA bases to specify each amino acid. (This reasoning assumes that each amino acid is encoded by the same size block of RNA.)
In addition, a ribosome must know where to start synthesizing a protein on an mRNA molecule and where to stop, and start and stop signals require their own RNA sequences. A series of experiments carried out in the 1960s confirmed these mathematical speculations, and went on to determine which triplet sequence (called a codon ) specifies which amino acid.
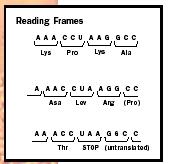
Indeed, the genetic code uses codons of three bases each, such as ACC or CUG. Therefore, the protein synthesis machinery reads every triplet of bases along the mRNA and builds a chain of amino acids—a protein—accordingly. Reading triplets, however, would allow a ribosome to start at any one of three positions within a given triplet (see Fig. 1). The position that the ribosome chooses is based on the location of the start signal and is called the "reading frame."
Experiments have shown all but three of the sixty-four possible codons that A, G, U, and C specify code for one amino acid each. This means that most amino acids are encoded by more than one codon. In other words, the genetic code is said to be redundant or degenerate. This redundancy allows the protein-synthesizing machinery of the cell to get by with less, as will be seen below. The three that don't, the "nonsense" codons, indicate the end of the protein-coding region of an mRNA, and are termed stop codons.
Starting, Stopping, and Making Protein
In any mRNA molecule, one codon always marks the beginning of a protein. That "start" codon is usually AUG in both eukaryotes and prokaryotes , although eukaryotes use GUG on rare occasions. AUG codes for the amino acid methionine. To start synthesizing at an AUG, however, ribosomes require more information besides a start codon; this information is found in the sequence surrounding the initial AUG. AUG codons in the middle of a protein-coding sequence are translated like any other codon.
Three codons signal the end of the mRNA template. These so-called stop codons, UAA, UAG, and UGA, do not code for any amino acid. Instead, the ribosome gets stuck, waiting for the tRNA that never comes, and eventually falls off, releasing the newly synthesized protein.
The complementary sequence of a codon found on a tRNA molecule is called its anticodon. The tRNA molecule matches up its anticodon with the correct codon on the mRNA. A tRNA molecule holds an amino acid in one of its molecular arms and works with the ribosome to add its amino acid to the protein being synthesized. Each tRNA is then reloaded with its specific amino acid by an enzyme in the cytosol .
Codons that code for the same amino acid are called redundant codons. The first two bases of redundant codons are usually the same and the third is either U or C, or alternatively A or G. For example, two redundant codons for the amino acid arginine are CGU and CGC, both of which pair with the same tRNA, despite having different third bases. This characteristic of the codon-anticodon interaction is called "wobble," and it allows organisms to have fewer than sixty-four distinct tRNA genes. In some tRNAs, wobble is made possible by a modified base within the anticodon. This modified base is called inosine (designated by I) and is made from adenine.
Evidence for Evolution
For almost all organisms tested, including humans, flies, yeast, and bacteria, the same codons are used to code for the same amino acids. Therefore,

Mammalian mitochondria , which contain DNA, use the codon UGA not as a stop signal but instead to specify the amino acid tryptophan, and they have four stop codons instead of three. Also, the modified base inosine is not used in mitochondrial anticodons. Mitochondrial genetic codes from different organisms can also be distinct from each other as well as from the universal code, reflecting both their ancient bacterial origins and their long isolation within their host species.
SEE ALSO Archaea ; Cell Evolution ; DNA ; Eubacteria ; Gene ; Mitochondrion ; Nucleotides ; Protein Synthesis ; Protista ; Ribosome ; Transcription
Mary Beckman
Bibliography
Alberts, Bruce, et al. Molecular Biology of the Cell, 4th ed. New York: Garland Publishing, 2000.
Creighton, Thomas E. Proteins: Structures and Molecular Properties, 2nd ed. New York: W. H. Freeman, 1993.
Freifelder, David. Molecular Biology, 2nd ed. Boston: Jones & Bartlett, 1987.
Lehninger, Albert L. Principles of Biochemistry, New York: Worth Publishers, 1982.
Comment about this article, ask questions, or add new information about this topic: